The Meek Lab is investigating the population genomics of the ESA listed California Central Valley Chinook salmon (Oncorhynchus tshawytscha). We are using next-generation sequencing techniques (RAD-seq) to evaluate population structure within the Central Valley and to look for signatures of natural selection across the genome. Thus far, by taking a genome-wide approach, we’ve discovered over 12,000 new single nucleotide polymorphisms (SNPs) and show greater population structuring and genetic diversity in the Central Valley than previously identified (Meek et al. 2019, O’Leary et al. 2021). Additionally, we have developed an ancestry informative marker set of SNP markers that improve our ability to distinguish between the various Chinook salmon run types and identify unknown individuals back to their source population (Meek et al. 2016). This work improves our understanding of the genomic structure involved in adaptive variation among populations. It will be informative in the management of this imperiled species, including evaluating potential source populations for reintroduction efforts, genetic monitoring of populations in the face of climate change, and identifying juveniles to run type.
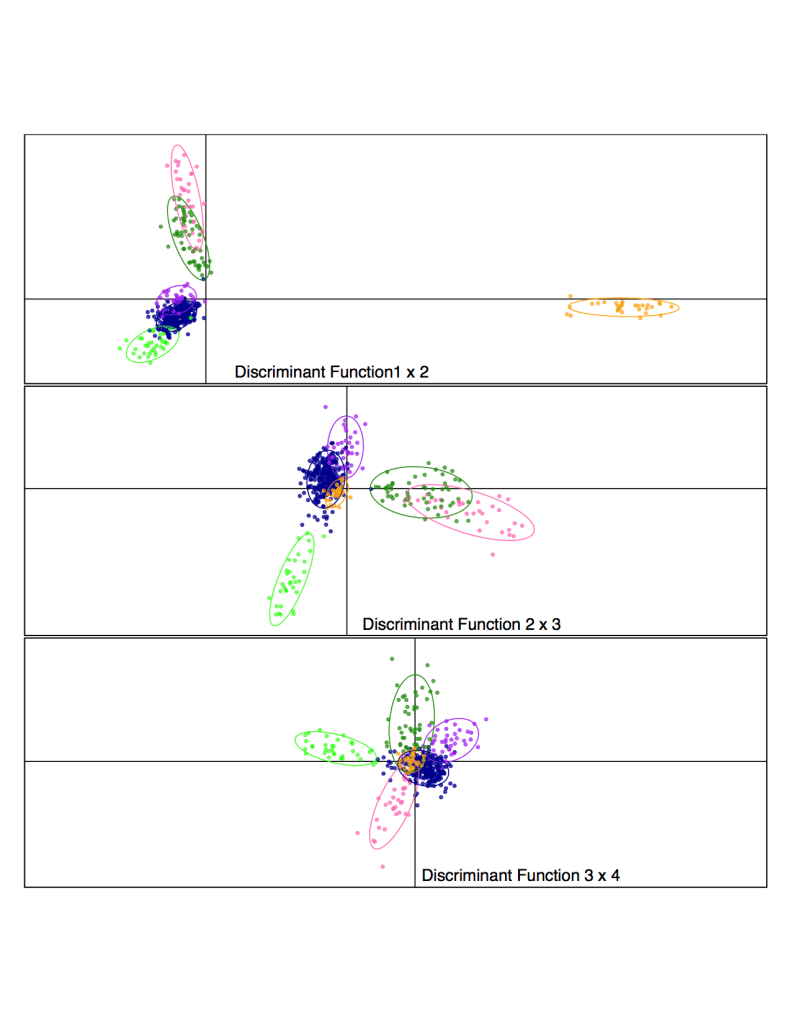
Population structure of Central Valley Chinook salmon shown by discriminant analysis of principal components, using ~12,000 discovered SNPs. Colors tightly correspond with run timing: Light green = Late Fall run, Blue = Fall run, Orange = Winter run, Purple = Spring from Feather River Hatchery, Dark green = Spring from Mill and Deer Crs., Pink = Spring from Butte Cr.
The Meek Lab has recently been funded by the Delta Science Program to continue this exciting work to improve our ability to protect life history trait diversity in Chinook salmon. Life history trait diversity (migration timing) is decreasing in the Central Valley, leading to increased demographic synchrony and decreased buffering. A key roadblock to protecting the diversity present is the current inability to rapidly and inexpensively identify large numbers of individuals from different populations and life history types during their outmigration. We are addressing this information gap by leveraging pre-existing genomic data to develop a new high-throughput genotyping panel that will allow us to identify individuals to run type and tributary, including identifying Fall run that are from the Sacramento versus the San Joaquin River basins. Using our newly developed molecular panel, we are genotyping thousands of juvenile Chinook salmon tissue samples collected over the past 20+ years. These individuals are being assigned to their run and tributary or river basin of origin. We are using this information, in combination with environmental data on water temperature and river flows, to determine the relationship between environmental conditions and yearly juvenile life history diversity. The information generated by this work will provide managers with the ability to accurately monitor the effect of key management actions on the different Central Valley Chinook salmon populations.

PRESS COVERAGE:
“Mariah Meek Harnesses the Power of New Genomic Tools to Address a Real-world Conservation Problem” https://integrativebiology.natsci.msu.edu/news/mariah-meek-harnesses-the-power-of-new-genomic-tools-to-address-a-real-world-conservation-problem/
“MSU researcher nets grant to investigate imperiled Chinook salmon in California’s Central Valley” https://natsci.msu.edu/news/msu-researcher-nets-grant-to-investigate-imperiled-chinook-salmon-in-californias-central-valley/

RELATED PUBLICATIONS:
O’Leary, S., T. Thompson, and M. Meek. 2021. Every cog and wheel: Unraveling biocomplexity at the genomic and phenotypic level in a population complex of Chinook salmon. bioRxiv. https://www.biorxiv.org/content/10.1101/2021.03.26.437213v1.full
Meek, M. Stephens, A. Goodbla, B. May, and M. Baerwald. 2019. Identifying hidden population structure and genomic diversity in Chinook salmon, a migratory species with a history of anthropogenic influence. Accepted at Canadian Journal of Fisheries and Aquatic Sciences. pdf
Meek, M., M. Baerwald, M. Stephens, A. Goodbla, K. Tomalty, M. Miller, and B. May. 2016. Sequencing improves our ability to study threatened migratory species: genetic population assignment in California’s Central Valley Chinook salmon. Ecology and Evolution. DOI: 10.1002/ece3.2493
Tomalty, K., M. Meek, M. Stephens, G. Rincón, N. Fangue, B. May, and M. Baerwald. 2015. Transcriptional response to acute thermal stress in juvenile Chinook salmon, Oncorhynchus tshawytscha, determined by RNAseq. G3: Genes, Genomes, Genetics. 5(7): 1335-1349 (pdf)
Meek, M., M. Stephens, A. Wong. K. Tomalty, B. May, M. Baerwald. 2014. Genetic characterization of California’s Central Valley Chinook salmon. Ecology. 95(5):1431.
Tomalty, K., M. Stephens, M. Baerwald, K. Bork, M. Meek, and B. May. 2014. Genetic considerations for fall-run Chinook salmon during the San Joaquin River Restoration. San Francisco Estuary and Watershed Science. 12(2): jmie_sfews_14880. Online at http://escholarship.org/uc/item/7bp9m8t9